Chinese Journal of Applied Entomology ›› 2020, Vol. 57 ›› Issue (4): 911-920.doi: 10.7679/j.issn.2095-1353.2020.093
• Research Articles • Previous Articles
Zheng-Hong CAO1,**( ), Kang HE1, Le XU1, Shen-Yang TANG1, Ya-Qin WANG2,3, Fei LI1,***(
), Kang HE1, Le XU1, Shen-Yang TANG1, Ya-Qin WANG2,3, Fei LI1,***( )
)
Received:2020-05-06
Accepted:2020-06-04
Online:2020-07-27
Published:2020-09-02
Contact:
Zheng-Hong CAO,Fei LI
E-mail:3130100574@zju.edu.cn;lifei18@zju.edu.cn
Zheng-Hong CAO, Kang HE, Le XU, Shen-Yang TANG, Ya-Qin WANG, Fei LI. Change in the gene expression of seedling stage rice in response to feeding by the brown planthopper Nilaparvata lugens (Stål)(Hemiptera: Delphacidae)[J]. Chinese Journal of Applied Entomology, 2020, 57(4): 911-920.
| 基因ID Gene ID | 上游引物Forward primer (5?-3?) | 下游引物Reverse primer (5?-3?) |
|---|---|---|
| Ubiquitin 5 (UBQ5) | AACCACTTCGACCGCCACT | GTTCGATTTCCTCCTCCTTCC |
| OsR498G0100081600 | CGACACCATGATCCGTCTCCC | CACTTCCACGGCCTCTTCG |
| OsR498G0100081800 | ACCATCCTGCTCTTCCTCCTCG | GGCGGGTTCATCTTGTTGC |
| OsR498G0102334700 | GAACGCAAGAATCCGCTCCC | TCCCAGCTTGTGCAGTCCC |
| OsR498G0203907000 | CGTCATGTGGGAGATGGAGCAG | CTGGTGTTGGTGGTGGTGGC |
| OsR498G0306783600 | AAACAACCATCCCGCTGCTA | TCATCCTTCCTCCGCCTC |
| OsR498G0408187800 | CATTGCCTCCGTTTCCTC | TTTCCAGCAGCAGCGATT |
| OsR498G0408202600 | ATGTCGGCGAGGATGAAC | CCCTAAGCAAGCGAAACC |
| OsR498G0408737000 | ACCGGCGACGACAGGGACAA | CCAGTAGGGAGGCCAGTAGCAC |
| OsR498G1018988800 | AGCGATACTGTGATTCGGTCAA | TGGCGGATGAGGATTTGG |
| OsR498G1019021500 | ATGGCTGACGCAGAGGAC | CCCAACCATAACGCCTGT |
| OsR498G1019092700 | GGTGATGTCCAGCCTCGGGTTC | GTTGGCGATGGCGAAGACG |
| OsR498G1120631700 | CACCATCAATCCATTCCTCCAT | CTCGCCATCCTGAACTGC |
| OsR498G1120633300 | GCACTTCTTCATGCACGACACG | GACCACCGACAGCTCCCTCA |
| OsR498G1221501200 | GGAAAGTGCTGCTCGGATGA | ATGCCCTTGCTGTTGTGG |
| OsR498G0100558200 | GCGGAGGCGAGGAACATCAAG | CATGGCTTCTTGGCCTGCTTGT |
| OsR498G0100612200 | TCAGCCGCCTCTTCTCCA | GCAACACCGCATCCCTCA |
| OsR498G0101116900 | GGGAGGAGTGGCGTGCTGAA | GGAAGGGACGAGGTTTACG |
| OsR498G0101439000 | TGATGATGATGGTTACGCAGAC | CACGGTGATAGCACAAACTCTT |
| OsR498G0203527800 | CACCGTCAACGTCACCACCG | TCTGCGGCTTCCCGTGCTTC |
| OsR498G0203880500 | AACGGCTTCGGTTTACGC | CAACTCCAAAGCGAAATGTAA |
Table 1 qPCR primers
| 基因ID Gene ID | 上游引物Forward primer (5?-3?) | 下游引物Reverse primer (5?-3?) |
|---|---|---|
| Ubiquitin 5 (UBQ5) | AACCACTTCGACCGCCACT | GTTCGATTTCCTCCTCCTTCC |
| OsR498G0100081600 | CGACACCATGATCCGTCTCCC | CACTTCCACGGCCTCTTCG |
| OsR498G0100081800 | ACCATCCTGCTCTTCCTCCTCG | GGCGGGTTCATCTTGTTGC |
| OsR498G0102334700 | GAACGCAAGAATCCGCTCCC | TCCCAGCTTGTGCAGTCCC |
| OsR498G0203907000 | CGTCATGTGGGAGATGGAGCAG | CTGGTGTTGGTGGTGGTGGC |
| OsR498G0306783600 | AAACAACCATCCCGCTGCTA | TCATCCTTCCTCCGCCTC |
| OsR498G0408187800 | CATTGCCTCCGTTTCCTC | TTTCCAGCAGCAGCGATT |
| OsR498G0408202600 | ATGTCGGCGAGGATGAAC | CCCTAAGCAAGCGAAACC |
| OsR498G0408737000 | ACCGGCGACGACAGGGACAA | CCAGTAGGGAGGCCAGTAGCAC |
| OsR498G1018988800 | AGCGATACTGTGATTCGGTCAA | TGGCGGATGAGGATTTGG |
| OsR498G1019021500 | ATGGCTGACGCAGAGGAC | CCCAACCATAACGCCTGT |
| OsR498G1019092700 | GGTGATGTCCAGCCTCGGGTTC | GTTGGCGATGGCGAAGACG |
| OsR498G1120631700 | CACCATCAATCCATTCCTCCAT | CTCGCCATCCTGAACTGC |
| OsR498G1120633300 | GCACTTCTTCATGCACGACACG | GACCACCGACAGCTCCCTCA |
| OsR498G1221501200 | GGAAAGTGCTGCTCGGATGA | ATGCCCTTGCTGTTGTGG |
| OsR498G0100558200 | GCGGAGGCGAGGAACATCAAG | CATGGCTTCTTGGCCTGCTTGT |
| OsR498G0100612200 | TCAGCCGCCTCTTCTCCA | GCAACACCGCATCCCTCA |
| OsR498G0101116900 | GGGAGGAGTGGCGTGCTGAA | GGAAGGGACGAGGTTTACG |
| OsR498G0101439000 | TGATGATGATGGTTACGCAGAC | CACGGTGATAGCACAAACTCTT |
| OsR498G0203527800 | CACCGTCAACGTCACCACCG | TCTGCGGCTTCCCGTGCTTC |
| OsR498G0203880500 | AACGGCTTCGGTTTACGC | CAACTCCAAAGCGAAATGTAA |
| 样品名称 Sample ID | Q30碱基比率 Q30 base ratio | 低质量读长比率 Low quality read ratio | 高质量reads High quality reads | 参考基因组比对率 Reference genome alignment rate (%) |
|---|---|---|---|---|
| CK1 | 96.15 | 0.81 | 60 034 869 | 93.6 |
| CK2 | 96.06 | 0.86 | 58 905 381 | 92.8 |
| CK3 | 96.05 | 0.77 | 57 745 824 | 92.6 |
| BFR1 | 96.31 | 0.62 | 53 269 890 | 93.4 |
| BFR2 | 95.94 | 0.85 | 56 389 786 | 92.4 |
| BFR3 | 96.02 | 0.75 | 63 845 983 | 92.1 |
Table 2 Transcriptome sequencing data quality assessment
| 样品名称 Sample ID | Q30碱基比率 Q30 base ratio | 低质量读长比率 Low quality read ratio | 高质量reads High quality reads | 参考基因组比对率 Reference genome alignment rate (%) |
|---|---|---|---|---|
| CK1 | 96.15 | 0.81 | 60 034 869 | 93.6 |
| CK2 | 96.06 | 0.86 | 58 905 381 | 92.8 |
| CK3 | 96.05 | 0.77 | 57 745 824 | 92.6 |
| BFR1 | 96.31 | 0.62 | 53 269 890 | 93.4 |
| BFR2 | 95.94 | 0.85 | 56 389 786 | 92.4 |
| BFR3 | 96.02 | 0.75 | 63 845 983 | 92.1 |
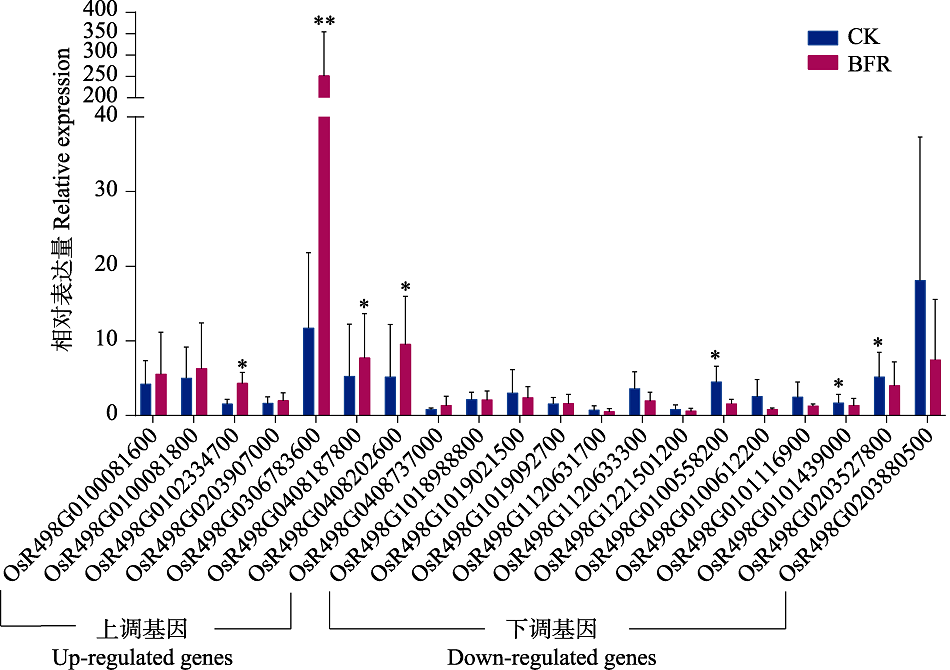
Fig. 2 The quantitative real-time PCR validation of 20 differentially expressed genes in rice seedling tissue induced by BPH feeding BFR: Rice samples after feeding by brown planthopper; CK: Rice samples in the control without feeding. Student’s T-test is used for statistical analysis, and data are mean ± standard errors (SE). * stands for significant difference (P < 0.05), ** stands for extremely significant difference (P < 0.01).
| 转录因子类型 Transcription factor type | 上调基因数目 Up-regulated gene number | 下调基因数目 Down-regulated gene number |
|---|---|---|
| MYB | 5 | 4 |
| ERF | 6 | 1 |
| ARF | 2 | 4 |
| NAC | 4 | 1 |
| HOX | 2 | 1 |
| Zc3h | 2 | 2 |
| WRKY | 6 | 0 |
| bHLH | 0 | 4 |
| Bzip | 1 | 1 |
| AHL | 2 | 0 |
| 其他Others | 6 | 7 |
| 总计Total number | 36 | 25 |
Table 3 The differentially expressed transcription factors in rice seedling tissue induced by BPH feeding
| 转录因子类型 Transcription factor type | 上调基因数目 Up-regulated gene number | 下调基因数目 Down-regulated gene number |
|---|---|---|
| MYB | 5 | 4 |
| ERF | 6 | 1 |
| ARF | 2 | 4 |
| NAC | 4 | 1 |
| HOX | 2 | 1 |
| Zc3h | 2 | 2 |
| WRKY | 6 | 0 |
| bHLH | 0 | 4 |
| Bzip | 1 | 1 |
| AHL | 2 | 0 |
| 其他Others | 6 | 7 |
| 总计Total number | 36 | 25 |
| 类型Type | 基因编号Gene ID | 基因注释Gene annotation | 差异倍数 Fold change |
|---|---|---|---|
| SA相关基因 Genes associated with SA | OsR498G0815887900 | Protein SAR DEFICIENT 1 | 3.76 |
| OsR498G0100620200 | Transcription factor WRKY33 | 2.86 | |
| OsR498G0511031400 | Probable WRKY transcription factor 33 | 2.85 | |
| OsR498G1221518200 | Alpha-dioxygenase 1(DOX1) | 2.13 | |
| JA相关基因 Genes associated with JA | OsR498G1221890200 | Transcription factor JAMYB | 2.88 |
| OsR498G0204402500 | Transcription factor MYB21 | 3.79 | |
| OsR498G0100288900 | Probable WRKY transcription factor 72 | 2.19 | |
| OsR498G0816439800 | Lipoxygenase 7, chloroplastic(LOX7) | 3.39 | |
| OsR498G0306871300 | Lipoxygenase-1(LOX1) | 4.27 | |
| OsR498G0509800400 | Acyl-coenzyme A oxidase 4, peroxisomal(ACX4) | 2.14 | |
| OsR498G0100209800 | Acyl-coenzyme A oxidase 4, peroxisomal(ACX4) | 2.00 | |
| OsR498G0510943100 | Cytochrome P450 94C1(CYP94C1) | 2.28 | |
| SA和JA相关基因 Genes associated with SA and JA | OsR498G1119325200 | Probable WRKY transcription factor 70 | 2.27 |
| OsR498G0203096300 | WRKY transcription factor WRKY71 | 2.46 | |
| OsR498G0714720100 | Putative methylesterase 11, chloroplast(MES11) | 2.28 |
Table 4 The differently expressed genes associated with SA and JA in rice seedling tissue induced by BPH feeding
| 类型Type | 基因编号Gene ID | 基因注释Gene annotation | 差异倍数 Fold change |
|---|---|---|---|
| SA相关基因 Genes associated with SA | OsR498G0815887900 | Protein SAR DEFICIENT 1 | 3.76 |
| OsR498G0100620200 | Transcription factor WRKY33 | 2.86 | |
| OsR498G0511031400 | Probable WRKY transcription factor 33 | 2.85 | |
| OsR498G1221518200 | Alpha-dioxygenase 1(DOX1) | 2.13 | |
| JA相关基因 Genes associated with JA | OsR498G1221890200 | Transcription factor JAMYB | 2.88 |
| OsR498G0204402500 | Transcription factor MYB21 | 3.79 | |
| OsR498G0100288900 | Probable WRKY transcription factor 72 | 2.19 | |
| OsR498G0816439800 | Lipoxygenase 7, chloroplastic(LOX7) | 3.39 | |
| OsR498G0306871300 | Lipoxygenase-1(LOX1) | 4.27 | |
| OsR498G0509800400 | Acyl-coenzyme A oxidase 4, peroxisomal(ACX4) | 2.14 | |
| OsR498G0100209800 | Acyl-coenzyme A oxidase 4, peroxisomal(ACX4) | 2.00 | |
| OsR498G0510943100 | Cytochrome P450 94C1(CYP94C1) | 2.28 | |
| SA和JA相关基因 Genes associated with SA and JA | OsR498G1119325200 | Probable WRKY transcription factor 70 | 2.27 |
| OsR498G0203096300 | WRKY transcription factor WRKY71 | 2.46 | |
| OsR498G0714720100 | Putative methylesterase 11, chloroplast(MES11) | 2.28 |
| [1] |
Castillo-Davis CI, Hartl DL , 2003. GeneMerge-post-genomic analysis, data mining, and hypothesis testing. Bioinformatics, 19(7):891-892.
doi: 10.1093/bioinformatics/btg114 URL pmid: 12724301 |
| [2] | Diao YH, Sun KD, Zhang YY, Wang XK , 2019. Research progress on drug resistance of brown planthopper and its control strategies. Agricultural Development and Equipment,(6): 28- 29, 34. |
| [ 刁跃珲, 孙凯迪, 张蕴邺, 王兴科 , 2019. 水稻褐飞虱抗药性研究进展及其治理对策. 农业开发与装备, (6): 28- 29, 34.] | |
| [3] |
Du HL, Yu Y, Ma YF, Gao Q, Cao YH, Chen Z, Ma B, Qi M, Li Y, Zhao XF, Wang J, Liu KF, Qin P, Yang X, Zhu LH, Li SG, Liang CZ , 2017. Sequencing and de novo assembly of a near complete indica rice genome. Nature Communications, 8:15324.
doi: 10.1038/ncomms15324 URL pmid: 28469237 |
| [4] |
Guo HM, Li HC, Zhou SR, Xue HW, Miao XX , 2014. Cis-12-oxo-phytodienoic acid stimulates rice defense response to a piercing-sucking insect. Molecular Plant, 7(11):1683-1692.
doi: 10.1093/mp/ssu098 URL |
| [5] |
Hu J, Xiao C, He YQ , 2016. Recent progress on the genetics and molecular breeding of brown planthopper resistance in rice. Rice, 9:30.
doi: 10.1186/s12284-016-0099-0 URL pmid: 27300326 |
| [6] | Ji D, Xin SJ , 2009. Development and data analysis of real-time fluorescent quantitative PCR. Letters in Biotechnology, 20(4):198-600. |
| [ 纪冬, 辛绍杰 , 2009. 实时荧光定量 PCR的发展和数据分析. 生物技术通讯, 20(4):598-600.] | |
| [7] |
Jing SL, Zhao Y, Du B, Chen RZ, Zhu LL, He GC , 2017. Genomics of interaction between the brown planthopper and rice. Current Opinion in Insect Science, 19(SI):82-87.
doi: 10.1016/j.cois.2017.03.005 URL |
| [8] |
Koo SC, Moon BC, Kim JK, Kim CY, Sung SJ, Kim MC, Cho MJ, Cheong YH , 2009. OsBWMK1 mediates SA-dependent defense responses by activating the transcription factor OsWRKY33. Biochemical and Biophysical Research Communications, 387(2):365-370.
doi: 10.1016/j.bbrc.2009.07.026 URL pmid: 19607808 |
| [9] |
Li B, Colin ND , 2011. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics, 12(1):1-16.
doi: 10.1186/1471-2105-12-1 URL |
| [10] |
Li CY, Luo C, Zhou ZH, Wang R, Ling F, Xiao LT, Lin YJ, Chen H , 2017. Gene expression and plant hormone levels in two contrasting rice genotypes responding to brown planthopper infestation. BMC Plant Biology, 17:57.
doi: 10.1186/s12870-017-1005-7 URL pmid: 28245796 |
| [11] |
Li R, Li JC, Zhou GX, Lou YG , 2013. Selection of internal reference genes in real-time quantitative PCR of rice pest-induced related genes. Chinese Bulletin of Botany, 48(2):184-191.
doi: 10.3724/SP.J.1259.2013.00184 URL |
| [ 李冉, 李建彩, 周国鑫, 娄永根 , 2013. 水稻虫害诱导相关基因实时定量PCR中内参基因的选择. 植物学报, 48(2):184-191.] | |
| [12] |
Lu HP, Luo T, Fu HW, Wang L, Tan YY, Huang JZ, Wang Q, Ye GY, Angharad MRG, Lou YG, Shu QY , 2018. Resistance of rice to insect pests mediated by suppression of serotonin biosynthesis. Nature Plants, 4(6):338-344.
doi: 10.1038/s41477-018-0152-7 URL pmid: 29735983 |
| [13] |
Matthias E, Philippe R , 2019. Molecular interactions between plants and insect herbivores. Annual Review of Plant Biology, 70(1):527-557.
doi: 10.1146/annurev-arplant-050718-095910 URL |
| [14] | Michael IL, Simon A, Wolfgang H , 2013. Differential analysis of count data-the DESeq2 package. http://www.bioconductor.org/packages/2.13/bioc/vignettes/deseq2/inst/doc/des.pdf. |
| [15] |
Vivek V, Pratibha R, Prakash PK , 2016. Plant hormone-mediated regulation of stress responses. BMC Plant Biology, 16:86.
URL pmid: 27079791 |
| [16] |
Yang L, Li A, Zhang WL , 2019. Current understanding of the molecular players involved in resistance to rice planthoppers. Pest Management Science, 75(10):2566-2574.
URL pmid: 31095858 |
| [17] |
Zhang HJ, Hong YB, Huang L, Liu SX, Tian LM, Dai Y, Cao ZY, Huang LH, Li DY, Song FM , 2016. Virus-induced gene silencing-based functional analyses revealed the involvement of several putative trehalose-6-phosphate synthase/phosphatase genes in disease resistance against Botrytis cinerea and Pseudomonas syringae pv. tomato DC3000 in tomato. Frontiers in Plant Science, 7:1176.
doi: 10.3389/fpls.2016.01176 URL pmid: 27540389 |
| [1] | Li-Tong SUN, Ling FENG, Zi-Rui LIU, Xiao-Wei XU, Jing-Lan LIU. Effects of trehalose on the physiological and biochemical characteristics of rice, including resistance to the brown planthopper (BPH), Nilaparvata lugens (Stål) (Hemiptera: Delphacidae) [J]. Chinese Journal of Applied Entomology, 2020, 57(4): 814-822. |
| [2] | Xue-Mei LI, Xiao-Xu ZHENG, Shuai-Jie HE, Feng-Lian YANG, Gang WU. Effects of different farming technologies on the population dynamics of pests and their natural enemies in rice fields [J]. Chinese Journal of Applied Entomology, 2020, 57(1): 105-112. |
| [3] | Xiao-Xu ZHENG, Mu-Hua ZHAO, Shuai-Jie HE, Xue-Mei LI, Feng-Lian YANG, Gang WU. Effects of five different rice varieties on the life history and population dynamics of the brown planthopper, Nilaparvata lugens in central China [J]. Chinese Journal of Applied Entomology, 2020, 57(1): 142-152. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
